Get purePlex™️ DNA
library prep system with
4 index sets
Featuring speed, performance, and auto-normalization.
seqWell Technology is Accelerating
NGS Research for Our Customers
Meet purePlex™ DNA Library Prep Kit
Maximize Your Sequencing Efforts
-
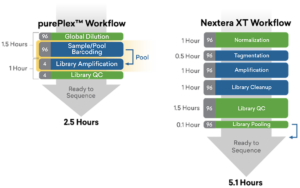
2.5-hour workflow for 96 samples, 45 min. hands-on time
-
Auto-normalization of read counts and insert size over 10-fold input range
-
Unique indices included
-
Significant cost and plastics savings over other transposase-based systems

SKU: 301067 | $2,015.00 USD
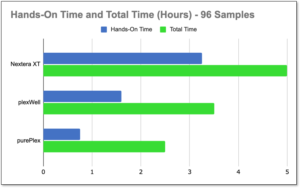
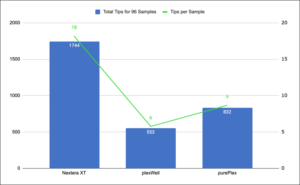
Fast, Easy Workflow with purePlex™
-
purePlex saves 50% total time over Nextera XT
-
75% hands-on time savings
-
All times based on processing of 96 samples
Learn more about how purePlex technology can maximize your sequencing efforts.
Save over 50% of time by using purePlex. Check out the workflow comparison
Saving $$$ and The Planet Through Less Plastic
-
50% + savings in plastics per sample
-
$7+ per plate savings
Complete the form to be contacted by a seqWell representative.
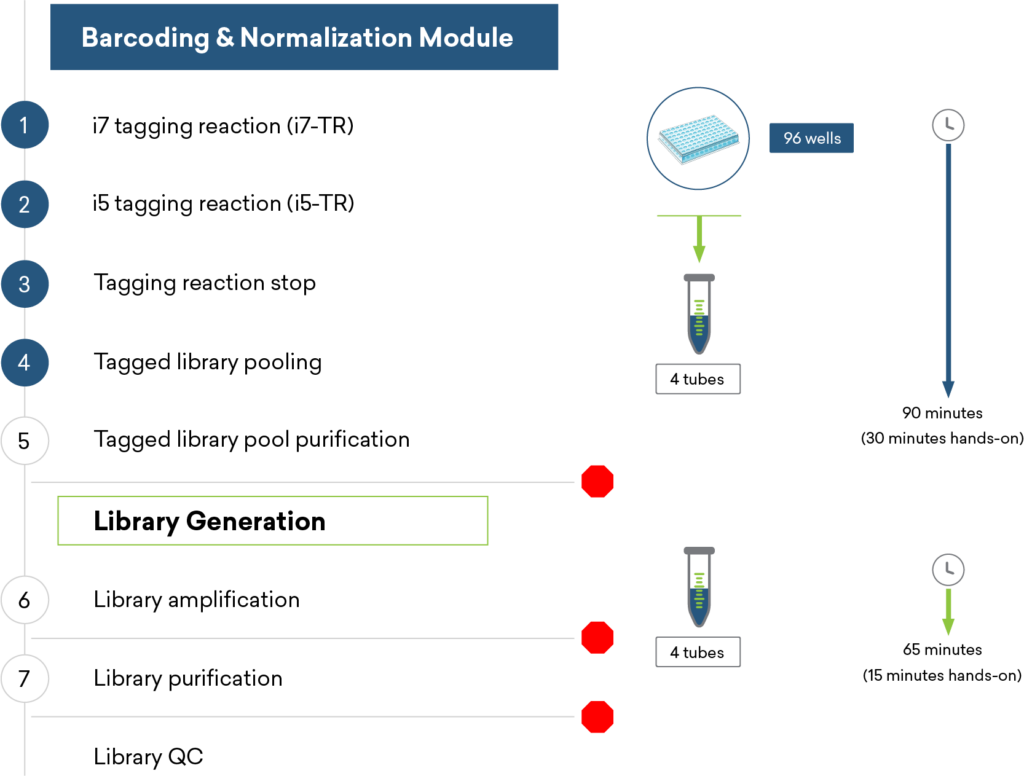
Benefits of purePlex:
-
Fast, flexible workflow with no requirement for full plate processing
-
Auto-normalization reduces QC burden, improves data consistency
-
Early pooling for easier sample handling
-
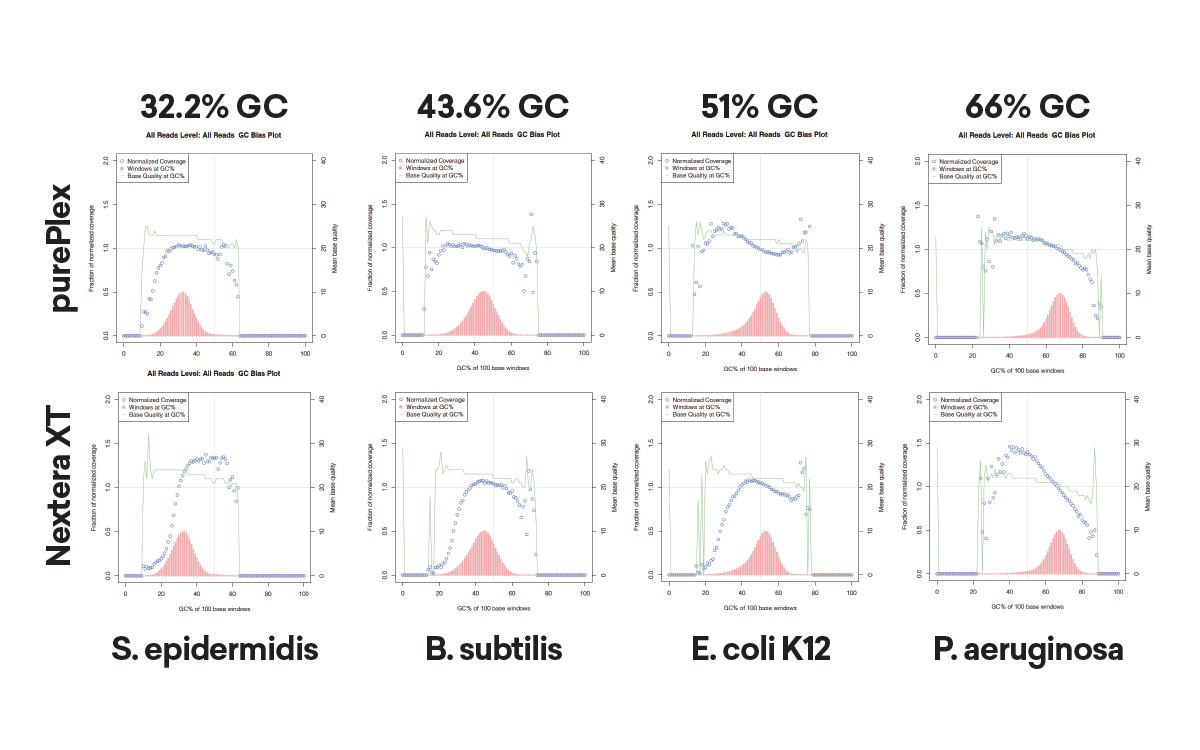
Reduced GC bias compared to other transposase-based methods
-
Significant cost and plastics savings
Product Highlights
Auto-normalization of insert size and read depth:
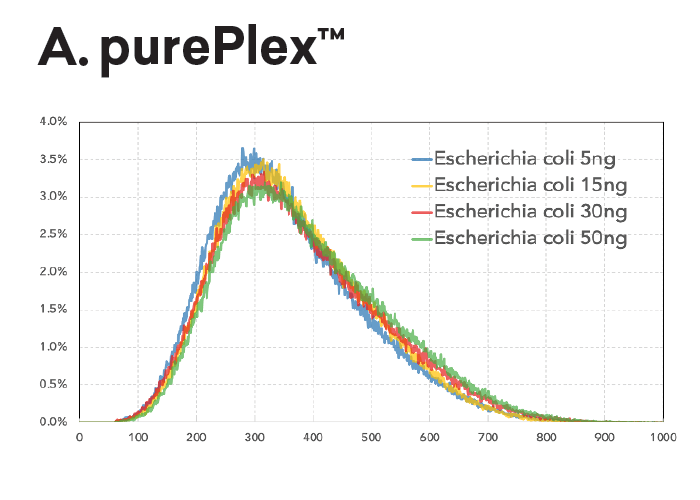
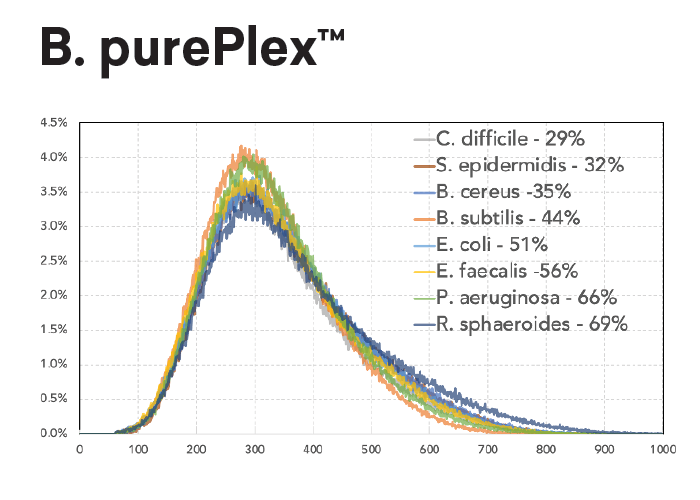
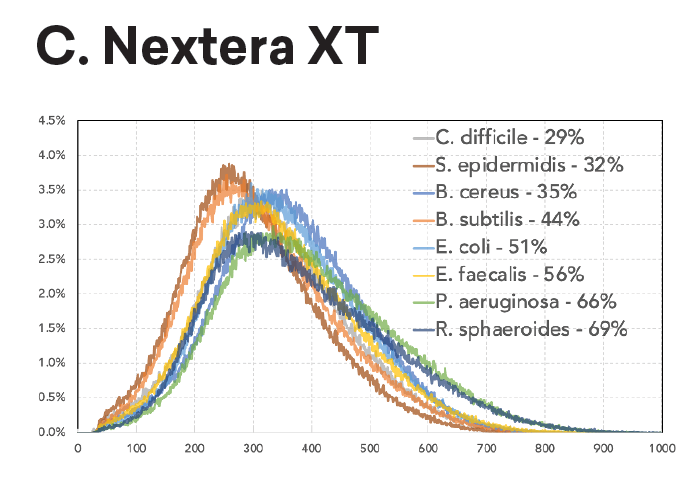
The insert size within a pool of samples is consistent regardless of input (panel A, left) or GC content (panel B, lower left). In contrast, Nextera XT libraries have varied fragment distributions from GC content even after normalizing the sample input.
Auto-normalization of Read Depth
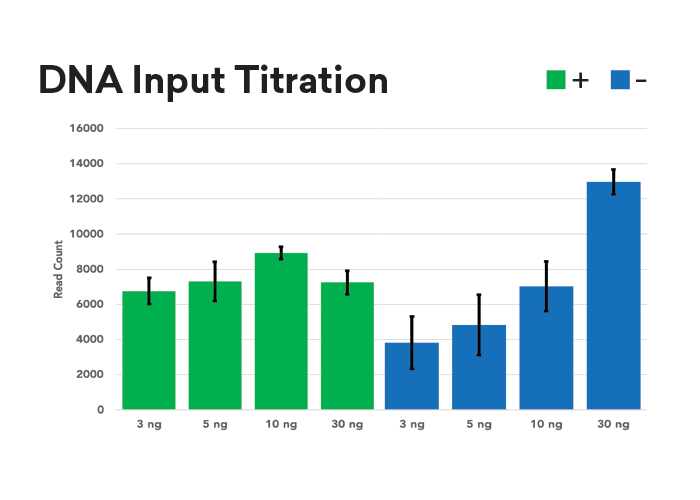
Samples were normalized to inputs of 3, 5, 10, and 30 ng then underwent purePlex library prep with (+) and without (–) normalization reagent. Read counts for each sample are equal, regardless of input, when normalization reagent is used. In contrast, without normalization reagent, sample read count scales with input.
Specifications
| Specs | Description |
|---|---|
| Sample Type | Amplicons, Plasmids, Genomic DNA, cDNA |
| DNA Input Range | 5 – 50 ng |
| Number of Unique Index Combinations | 2,304 |
| Supported Paired Reads (Clusters)/Sample | ≤ 20 million |
| Output Fragment Range | 400 – 1,200 bp |
| Applications | Synthetic construct sequencing (amplicons, plasmids, etc.), Low-pass whole genome sequencing, Whole small genome sequencing (<50 Mb), scRNA-seq, Metagenomics/Microbiome screening |
| Reactions per Kit | 96 |
purePlex DNA Library Prep Kit Includes:
-
i7 Tagging Reagent Plate
-
i5 Tagging Reagent Plate
-
Coding Buffer (3X)
-
X Solution
-
MAGwise Paramagnetic Beads
-
Normalization Reagent
-
Library Primer Mix