Discover true
multiplexing with
plexWell™ technology
Save time & cost with true multiplexed NGS libraries
seqWell Technology is Accelerating
NGS Research for Our Customers
Maximize Your Sequencing Efforts
Built-in Normalization = More Cost-effective Sequencing
-
Simplify your workflow by multiplexing up to 2,304 samples without the need for automation.
-
Skip time consuming normalization steps with built-in auto-normalization.
-
Save money and reduce waste with fewer tips required.

Perfect for your sequencing needs:
-
Whole microbial and metagenomic sequencing
-
Plasmids and PCR amplicons
-
SARS-CoV-2 sequencing
-
Low pass WGS
-
Single cell RNA-seq
Learn more about how plexWell technology can maximize your sequencing efforts.
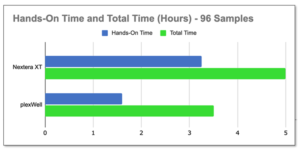
Save over 40% of time by using plexWell. Check out the workflow comparison
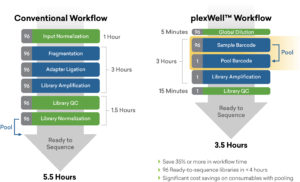
plexWell = Better Performance
plexWell™ Achieves a Significantly Better Level of Multiplexing Uniformity for Highly Multiplex Sequencing Applications.
Sequencing results obtained for sequencing 192 samples of amplified single-cell cDNA with plexWell™ (blue) and For Nextera™ reagents (orange). Input DNA was pre-normalized, whereas plexWell™ library was made from un-normalized amplified cDNA. Read count variation for plexWell™ showed 27% CV versus 71% CV for Nextera™.
Complete the form to be contacted by a seqWell representative.
Benefits of plexWell:
-
A “True Multiplexing” library prep platform technology for making normalized NGS libraries quickly and easily from large numbers of samples
-
Allows multiplexed libraries to be prepared as a single pool after a simple molecular tagging step
-
Normalizes across a wide range of input DNA amounts through chemistry
-
Transposon mediated
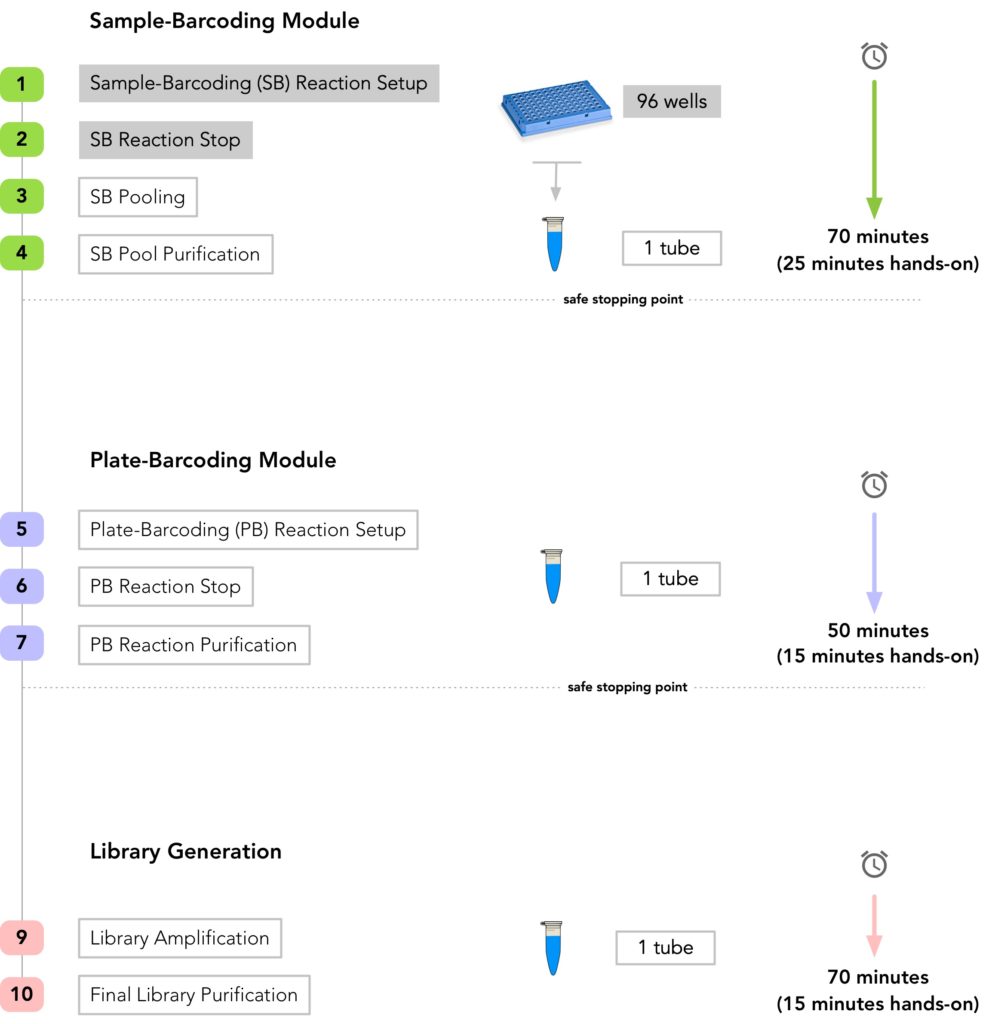
Simple & Scalable Workflow
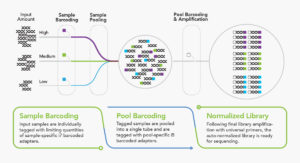
Integrated Normalization = More Samples Per Sequencing Run
Read Count Normalization Variable Input
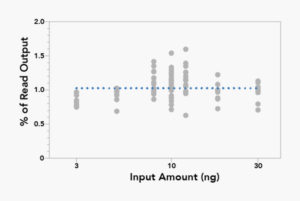
Compression of Input DNA Range
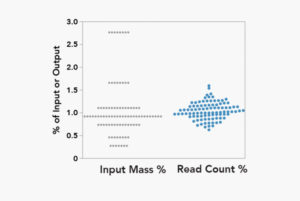
Precision Multiplexing
plexWell technology yields balanced multiplexed libraries containing highly uniform insert size distributions and samples read counts.
Median Insert Size Distribution for 96-plex Library:
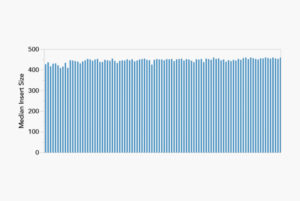
plexWell™ Insert Size Distribution:
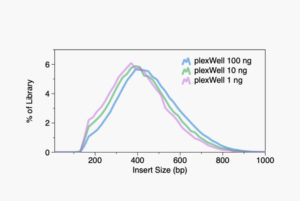
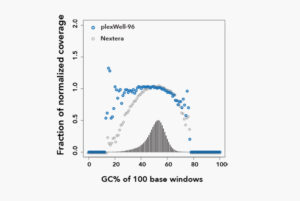
plexWell™ Delivers Minimal Bias for Diverse Applications
Microbial Genome
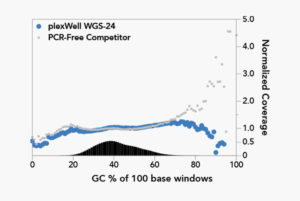
Mammalian Genome
Check out the plexWell workflow in action:
plexWell™ Library Prep Kits
plexWell™ library prep kits are available for all your sequencing applications. Feel free to check our plexWell™ products chart.