Addressing Plasmid QC with Quantitative Contamination Detection
Next-generation sequencing (NGS) plays a crucial role in various stages of therapeutic antibody development, from discovery and engineering to manufacturing. It enables the identification of high-affinity, highly specific candidates, optimization of expression systems, and quality control to maintain the genetic integrity, consistency, and safety of the therapeutic. One important consideration in antibody development is ensuring clonality, particularly when working with bacterial expression systems where co-transformation (Figure 1) can introduce variability1.
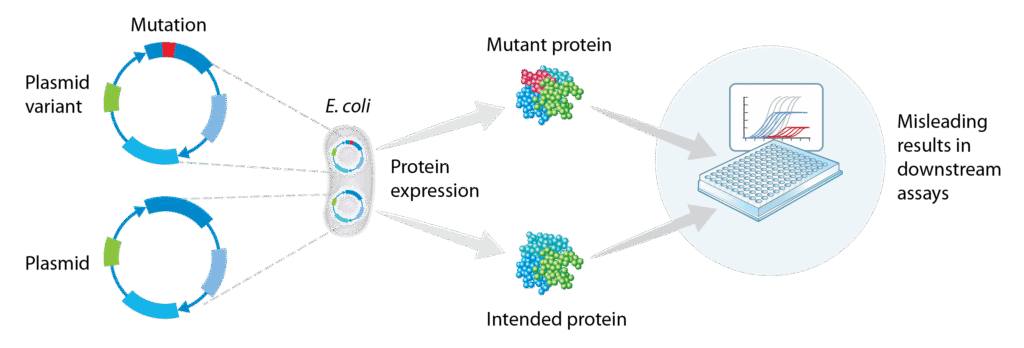
Figure 1. A single bacterial cell can host multiple plasmids resulting in impurities that can impact the results of downstream applications.
APPLICATION NOTE: Advances in Plasmid Verification- The Role of NGS in Identity & Clonality Testing.
Choosing the Right NGS Library Prep to Ensure Uninterrupted Antibody Development Pipelines
As sequencing technologies have evolved, the demand for higher sensitivity combined with higher throughput has increased in part to provide the ability to detect contaminating plasmids at scale. NGS can help address clonality concerns, providing confidence in sequence fidelity without creating a bottleneck within the antibody development process.
Due to the scale of synthetic construct usage within antibody development pipelines, both sequencing sensitivity and throughput are critical factors to consider. The choice of library prep method can greatly impact the reliability and overall throughput of generating NGS data, but is often overlooked. By leveraging a high-performance library preparation workflow that easily scales, NGS can help address these concerns, providing confidence in sequence accuracy and antibody purity without creating a bottleneck within the antibody development process.
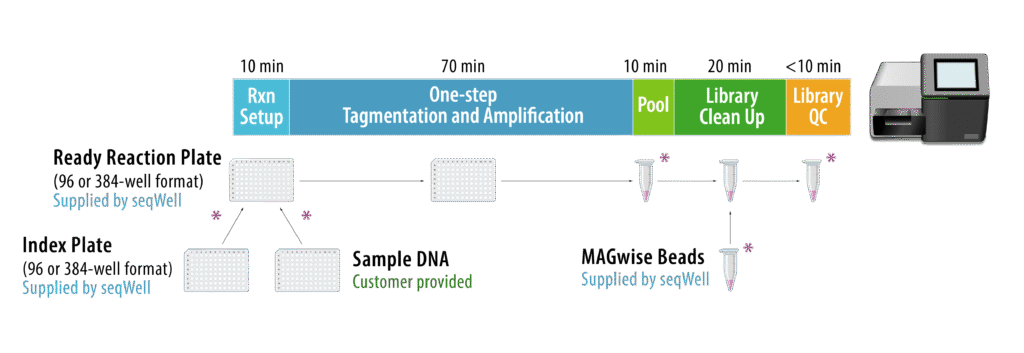
seqWell’s tagmentation-based, one-step ExpressPlexTM 2.0 Library Prep Kit is powered by TnX, the next generation transposase, to reduce insertional bias and improve sequence coverage uniformity using a highly streamlined workflow (Figure 2) for sequencing synthetic constructs like plasmids & amplicons.
Figure 2. ExpressPlex 2.0 library preparation kits utilize a proprietary mixture of enzymes to tag input DNA with indexed adapters and amplify libraries all in a single reaction. A 96-plex library can be prepared for library QC and sequencing in 100 minutes starting with plasmids or amplicons, with 30 minutes of hands-on time.
ExpressPlex 2.0 has demonstrated ultra-high throughput scalability with up to 6,144 indexes, a 384-well format, and wide automation compatibility that provides the capacity to process 1536 extracted DNA samples into a sequence-ready library in under 2.5 hours using nested processing2.
Competitor Tn5-transposase based tagmentation workflows are significantly more complex and less automation-friendly and require from 3 to 6 hours – depending on product – to process 96 samples from extracted DNA to sequence-ready libraries. These differences represent significant throughput limitations for construct screening.
POSTER: Echo Acoustic Liquid Handling and Next Generation TnX Transposase-based Technologies Enable Miniaturized, Automated NGS Library Preparation for Ultra High-Throughput Sequencing.
Read Count Normalization
The ExpressPlex 2.0 library prep kits combine fragmentation, barcoding and amplification into a single reaction, reducing hands‑on time to < 30 minutes and preparing a 96‑plex library in about 100 minutes. Built-in auto-normalization obviates the need to normalize sample input or output. The workflow generates a tight distribution of sequencing reads per sample (Figure 3), greatly reducing the QC burden upstream of highly multiplexed sequencing.
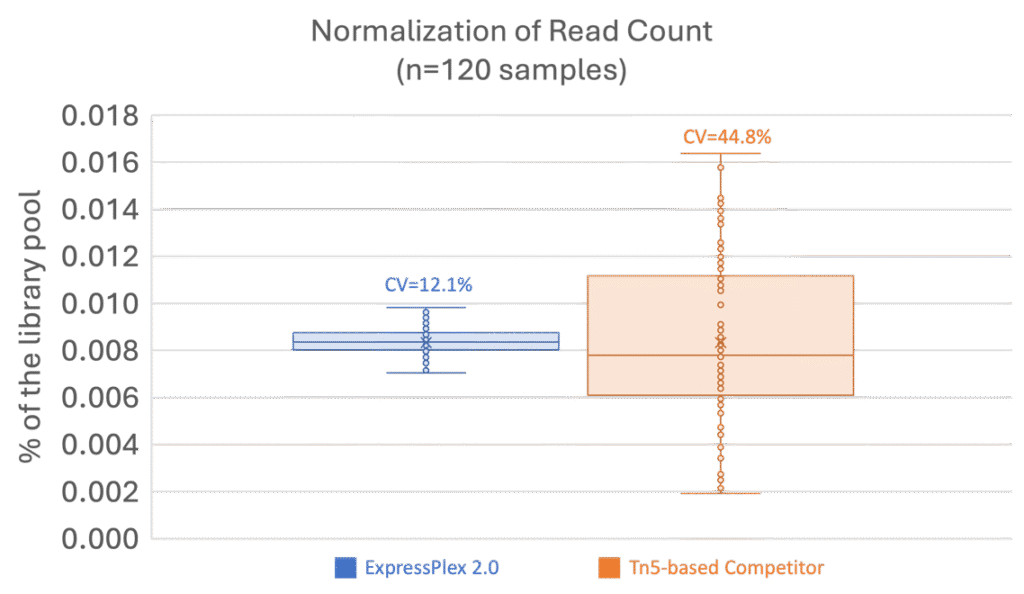
Figure 3. ExpressPlex 2.0 (blue) enables more consistent read count normalization with a c.v. of 12.1% across 120 samples, compared to a Tn5-transposase based competitor (orange) with a c.v. of 44.8% across the same number of samples.
Approach to Demonstrating the Sensitivity of the NGS/ExpressPlex 2.0 Workflow for Clonality Confirmation
Clonality concerns can be particularly pronounced in workflows relying on bacterial hosts, where co-transformation of multiple plasmids may occur, leading to contamination.
To simulate plasmid co-transformation, seqWell first designed a set of 5 high-quality plasmids with single-base mutations, addressing insertions, deletions, and substitutions (Figure 4). Plasmid concentrations were normalized and adjusted to 5.0 ng/µl based on fluorometric dsDNA assays. Additional experimental details are available3.

Figure 4. Plasmids designed with single-base mutations, including single-base substitutions from Adenine (A) to Cytosine (C), Adenine to Guanine (G), Adenine to Thymine (T), a single-base deletion, and a single-base insertion.
Next, ExpressPlex 2.0’s ability to detect co-transformation was investigated by spiking-in the plasmids carrying single-base mutations into a control plasmid background at 8 different levels: 0% (as a control), 0.1%, 0.2%, 0.4%, 0.5%, 1%, 10%, and 20%. Each spike-in level was run in triplicate to evaluate reproducibility.
A competitor’s Tn5 transposase-based kit was also used to prepare libraries from the same spike-in mixtures. The ExpressPlex and competitor’s libraries were pooled together prior to sequencing on the same run. A 2 x 150 bp run on the Illumina NextSeq 2000 (P1 flowcell) was used to sequence both the ExpressPlex and competitor libraries.
An acceptable industry-standard for the sensitivity assay was set at a level of 0.5% co-transformation variant (highlighted on Table 1).

Table 1. A comparison of expected vs. observed spike-in levels of plasmids carrying single-base mutations.
Table 1 shows performance equivalency between ExpressPlex 2.0 to an industry-standard Tn5 transposase-based library prep. ExpressPlex 2.0 generally follows the expected variant percentages, even at lower spike-in levels of below 1% (Figure 5), demonstrating its robust capability to detect plasmid co-transformation at various levels.
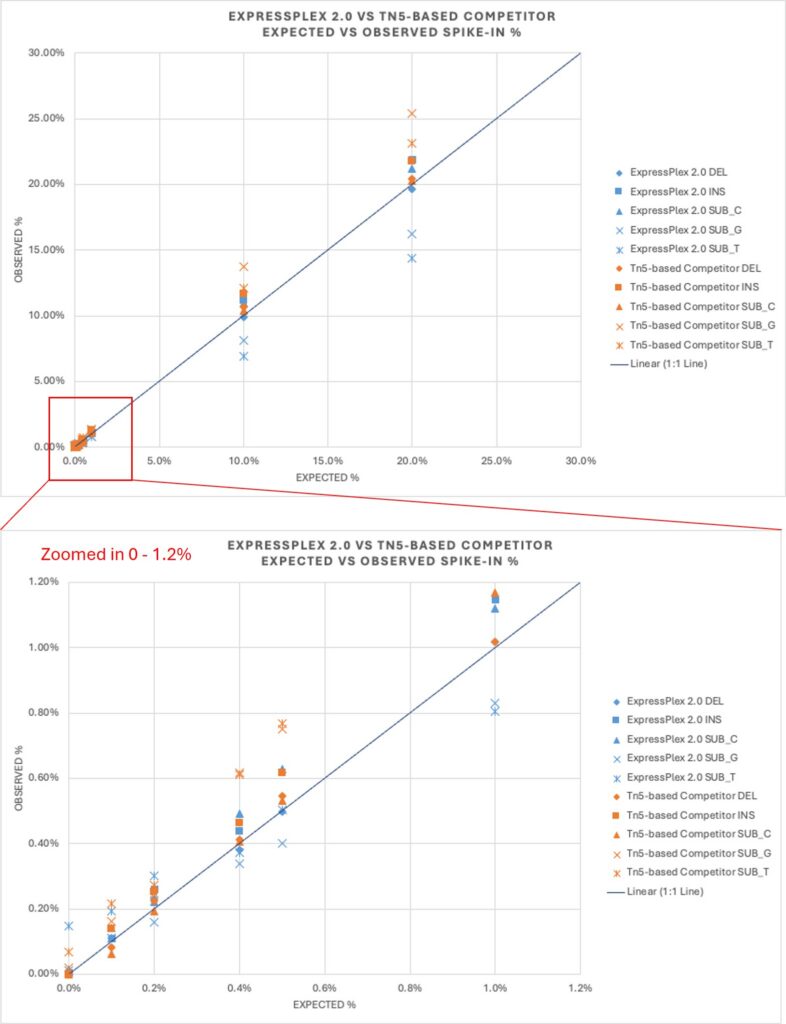
Figure 5. Accurately quantifying the presence of contaminating plasmids at levels below 1% demonstrates that ExpressPlex 2.0 is robust for quality control of plasmid purity.
Summary
Bacterial co-transformation presents a unique challenge, as the unintended presence of multiple plasmids within a single host cell can lead to spurious results. By utilizing a highly sensitive, easily scaled library preparation workflow, researchers can enhance their ability to detect and quantify low-level sequence variants, ensuring genetic homogeneity throughout the development process.
The ExpressPlex 2.0 Library Prep Kit offers a powerful and efficient solution to address these challenges. Its ability to rapidly and accurately identify plasmid co-transformation events makes it a well-suited tool for high-throughput plasmid quality control.
- ExpressPlex 2.0 Library Prep Kit: A highly sensitive, automation-friendly one-step workflow designed for sequence verification and assessing plasmid purity.
- High Sensitivity & Accuracy: Accurately quantifies contaminating plasmid differing only by a single basepair at levels below 1%, with high reproducibility across multiple spike-in levels.
- Enhanced Workflow Efficiency: 100-minute library prep with minimal hands-on time, built-in auto-normalization, and high-throughput scalability.
References:
- Tomoiaga D, Bubnell J, Herndon L, Feinstein P. High Rates of Plasmid Cotransformation in coli Overturn the Clonality Myth and Reveal Colony Development. Sci Rep. 2022 Jul 7; 12:11515.
- Echo Acoustic Liquid Handling and Next Generation TnX Transposase-based Technologies Enable Miniaturized, Automated NGS Library Preparation for Ultra High-Throughput Sequencing. Poster presented at the SLAS 2025 conference.
- Advances in Plasmid Verification: The Role of NGS in Identity & Clonality Testing. seqWell Application Note.