PhiRx™ Indexed Control
PhiRx Indexed Control is an easy-to-use solution to maximize the performance of XLEAP-SBS chemistry on high-throughput Illumina platforms by color-balancing both the sequencing reads and barcodes.
- Optimizes index diversity for XLEAP sequencing
- Serves as a seamless spike-in sequencing control, similar to Illumina’s PhiX Control V3
- Corrects index color balancing issues for both low diversity barcodes and small batch sequencing runs including 10x Genomics runs
- Improves signal uniformity and data quality
- Helps recover data that would be lost as undetermined reads
A Color-Balanced Spike-In Control for Improved Demultiplexing on XLEAP-SBS Illumina Sequencing Instruments
What is PhiRx?
PhiRx Indexed Control is a dual-indexed control library, made from phiX174 genomic DNA, optimized for Illumina sequencing platforms, particularly two-color systems like XLEAP SBS chemistry on NextSeq 1000/2000 and NovaSeq X or 10x Genomics runs. By improving color balancing for low-plexity libraries PhiRx ensures cleaner, more accurate sequencing results.
Key Features
- 10 bp i5 and i7 indexes with mixed bases: Provides high base diversity, automatically meeting color balancing requirements on supported platforms. (patent-pending)
- Color balancing optimization: Specifically designed for the sensitive XLEAP SBS chemistry, enabling as little as 5% spike-in to correct base calling issues in libraries with low-diversity indexes.
- High diversity for low-plexity samples: Randomized inserts with 45% GC content improve sequencing accuracy for low-diversity libraries.
- Reliable quality control: Derived from the well-characterized phiX174 genome, providing robust calibration to support sequencing accuracy and performance.
PhiRx Indexed Control reduces the rate of data loss from 53% to < 2%
| Spike-in Sequencing Control | % PhiX Aligned | % Undetermined Reads (Excluding PhiX) | % One-mismatch |
| seqWell PhiRx Indexed Control | 5.22 | 1.82 | 4.15 |
| Illumina PhiX Control V3 | 6.27 | 53.25 | 100.0 |
A 96-plex library prepared using ExpressPlex 2.0 was sequenced on a NextSeq 2000 with 5% spike-in of Illumina PhiX Control V3 or PhiRx from seqWell.
PhiRx: Data Recovery Case Study for 10x Genomics
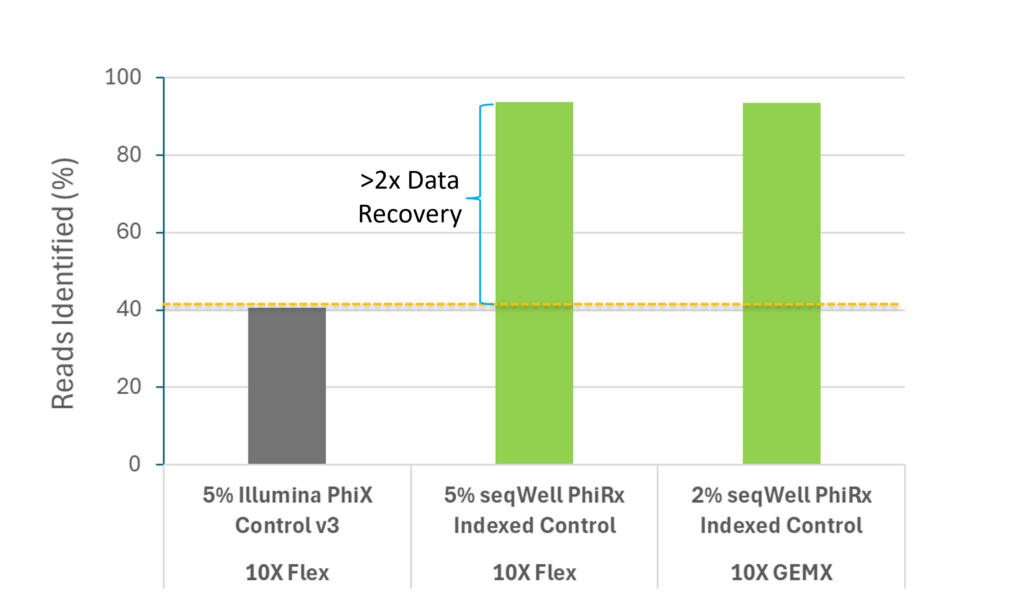
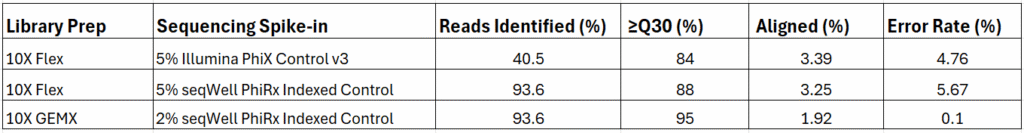
This data was generated from sequencing runs of 10X Flex and GEMX libraries with an Illumina PhiX Control V3 or with seqWell’s PhiRx Indexed Control equivalent spike-ins. The libraries were prepared by the Boston University Single Cell Sequencing Core and sequenced by the Microarray & Sequencing Resource.
“The Boston University Single Cell Sequencing Core accepts a variety of projects and some, like 10X Flex or GEMX, often require sequencing just one or a few multiplexed libraries at a time. Sequencing this on current XLEAP chemistry with Illumina unindexed PhiX would lose up to 80% of the data to unidentifiable reads due to low index complexity. By bioinformatically truncating the indexes or using just one of the two indexes, we can recover these reads but it takes additional time and effort and we are concerned about potential data misattribution in case of the index hopping. Sequencing accuracy and confidence in the data is important to us and our customers. We now happily use a 2-5% spike-in of the PhiRx Indexed Control which completely resolves this issue. “
– Yuriy Alekseyev, Ph.D. Director of the Single Cell Sequencing Core and the Microarray & Sequencing Resource at Boston University Medical Campus

Learn More in Our Blog

